Nat Med | Modus multi-omicus ad mappandum integratum tumoris, immunitatis et microbii prospectum cancri colorectali interactionem microbiomi cum systemate immuni revelat.
Quamquam indices biologici cancri coli primarii annis proximis late investigati sunt, praecepta clinica hodierna solum in stadiis metastasis tumoris-lymphonodi et detectione vitiorum reparationis discrepantiae DNA (MMR) vel instabilitatis microsatellitum (MSI) (praeter probationes pathologiae consuetas) nituntur ad commendationes curationis determinandas. Investigatores defectum associationis inter responsa immunia secundum expressionem geneticam, perfiles microbiales, et stroma tumoris in cohorte cancri colorectali Atlantis Genomatis Cancri (TCGA) et superviventiam aegrotorum animadverterunt.
Dum investigationes progrediuntur, notae quantitativae cancri colorectali primarii, inter quas natura cellularis, immunologica, stromalis, vel microbialis cancri, cum eventibus clinicis significanter conexae esse relatae sunt, sed adhuc limitata est intellectus quomodo interactiones earum eventus aegrotorum afficiant.
Ad nexum inter complexitatem phaenotypicam et exitum dissecandum, turma investigatorum ex Instituto Sidra Investigationis Medicae in Quataria nuper indicem integratum (mICRoScore) elaboravit et validavit qui coetum aegrotorum cum bonis ratibus supervivendi identificat, combinando proprietates microbiomi et constantes reiectionis immunis (ICR). Turma analysin genomicam comprehensivam exemplorum congelatorum recentium a 348 aegrotis cum cancro colorectali primario perfecit, inter quas sequentiatio RNA tumorum et textus colorectali sani congruentis, sequentiatio totius exomis, sequentiatio geni receptoris cellularum T profundarum et 16S rRNA bacterialis, supplementa sequentiatione genomae tumoris totius ad microbiome ulterius describendum. Studium in periodico "Nature Medicine" divulgatum est ut "Atlas integratus tumoris, immunitatis et microbiomis cancri coli".

Articulus in "Nature Medicine" editus.
AC-ICAM Summarium
Investigatores suggestum genomicum orthogonale adhibuerunt ad exempla tumorum congelatorum recentia analyzanda et ad tela coli sana adiacentia (paria tumoris normalis) ex aegris cum diagnosi histologica cancri coli sine therapia systemica comparanda. Fundamenta in sequentiatione totius exomis (WES), inspectione qualitatis datorum RNA-seq, et perscrutatione criteriorum inclusionis, data genomica ex 348 aegris retenta et ad analysin subsequentem cum mediana observationis 4.6 annorum adhibita sunt. Turma investigatorum hanc fontem "Sidra-LUMC AC-ICAM: Tabula et dux ad interactiones immune-cancer-microbiome" (Figura 1) nominavit.
Classificatio molecularis utens ICR
Capto modulari signorum geneticorum immunium ad continuam immunovigilantiam cancri, quod "constans immunis reiectionis" (ICR) appellatur, turma investigationis ICR optimizavit, condensando eum in tabulam viginti genorum, quae varia genera cancri, inter quae melanoma, cancer vesicae, et cancer mammae, comprehendit. ICR etiam cum responsione immunotherapiae in variis generibus cancri, incluso cancro mammae, coniunctum est.
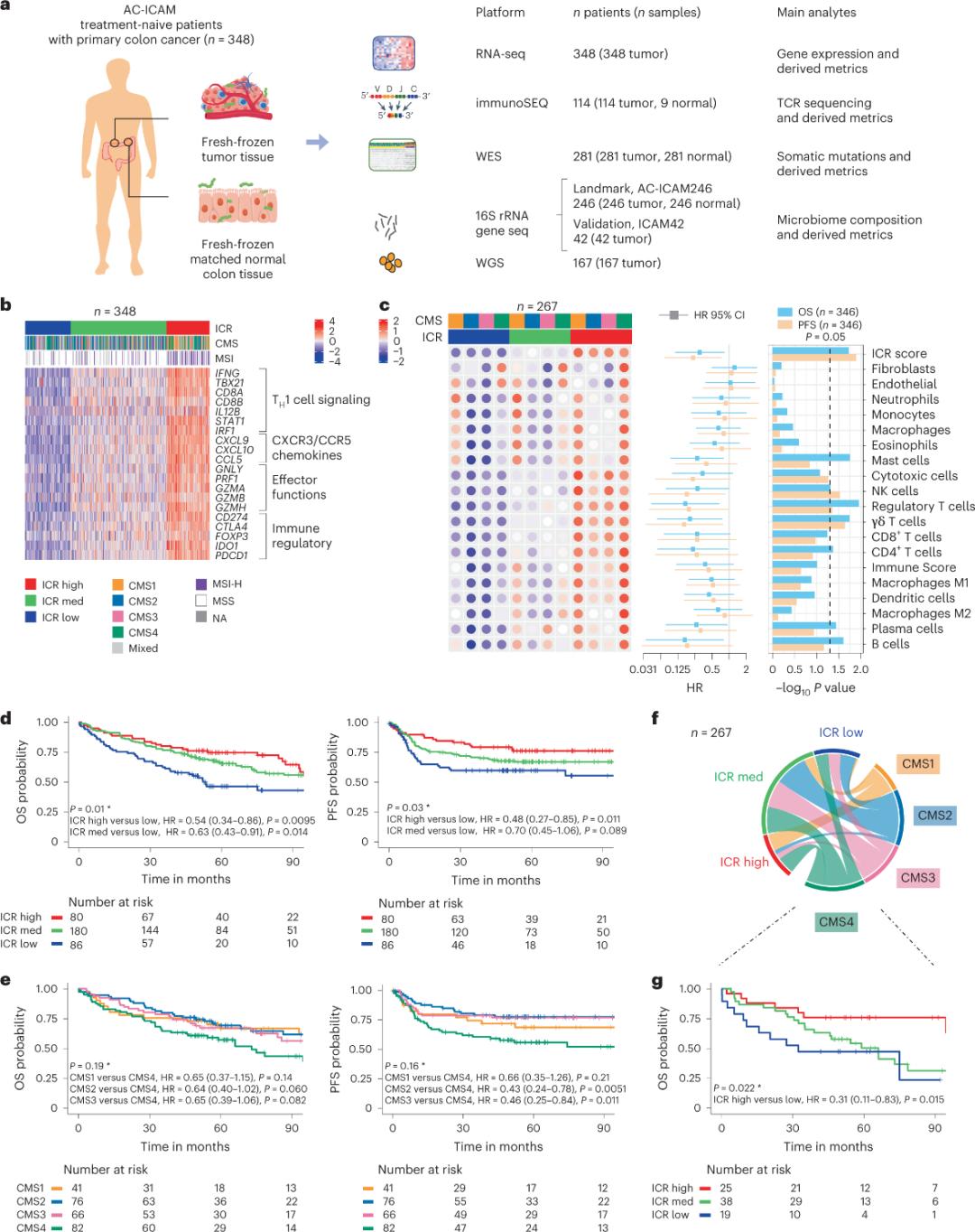
Primo, investigatores signaturam ICR cohortis AC-ICAM comprobaverunt, methodo co-classificationis secundum genes ICR utentes ad cohortem in tres greges/subtypos immunes classificandum: ICR altum (tumores calidos), ICR medium et ICR humilem (tumores frigidos) (Figura 1b). Investigatores propensionem immunem cum subtypis molecularibus consensualibus (CMS) associatam, classificationem cancri coli secundum transcriptoma, descripserunt. Categoriae CMS CMS1/immunem, CMS2/canonicum, CMS3/metabolicum et CMS4/mesenchymale includebant. Analysis ostendit indices ICR negative correlatos esse cum certis viis cellularum cancrariarum in omnibus subtypis CMS, et correlationes positivas cum viis immunosuppressivis et stromalibus conexis tantum in tumoribus CMS4 observatas esse.
In omnibus CMS, abundantia cellularum interfectricium naturalium (NK) et cellularum T maxima erat in subtypis ICR cum immunitate alta, cum variabilitate maiore in aliis subgroupis leucocytorum (Figura 1c). Subtypi immunitatis ICR differentes OS (Superviventiam Globalem) et PFS (Superviventiam Sine Progressu) habebant, cum incremento progressivo in ICR ab infimo ad altum (Figura 1d), quod munus prognosticum ICR in cancro colorectali validat.
Figura 1. Forma studii AC-ICAM, signatura genorum immuno-relata, subtypi immunis et molecularis, et superviventia.
ICR cellulas T tumore locupletatas et clonaliter amplificatas capit.
Pauca tantum cellularum T in tela tumoris infiltrantium specificae antigenis tumoris esse relatae sunt (minus quam 10%). Quapropter, maior pars cellularum T intra-tumoralium "cellulae T circumstantes" (cellulae T circumstantes) appellantur. Fortissima correlatio cum numero cellularum T conventionalium cum TCR productivis in subpopulationibus cellularum stromalium et leucocytorum observata est (per RNA-seq detecta), quae ad aestimandas subpopulationes cellularum T adhiberi potest (Figura 2a). In gregibus ICR (classificatio generalis et CMS), maxima clonalitas TCR immunium SEQ in gregibus ICR-high et CMS subtypi CMS1/immunibus observata est (Figura 2c), cum maxima proportione tumorum ICR-high. Usi toto transcriptoma (18,270 genis), sex gena ICR (IFNG, STAT1, IRF1, CCL5, GZMA, et CXCL10) inter decem gena optima cum clonalitate immunium SEQ TCR associata erant (Figura 2d). Clonalitas TCR ImmunoSEQ cum plerisque genis ICR fortius correlata est quam correlationes observatae utens signis CD8+ tumori respondentibus (Figura 2f et 2g). In conclusione, analysis supra suggerit signaturam ICR praesentiam cellularum T tumore locupletarum et clonaliter amplificatarum comprehendere et eius implicationes prognosticas explicare posse.
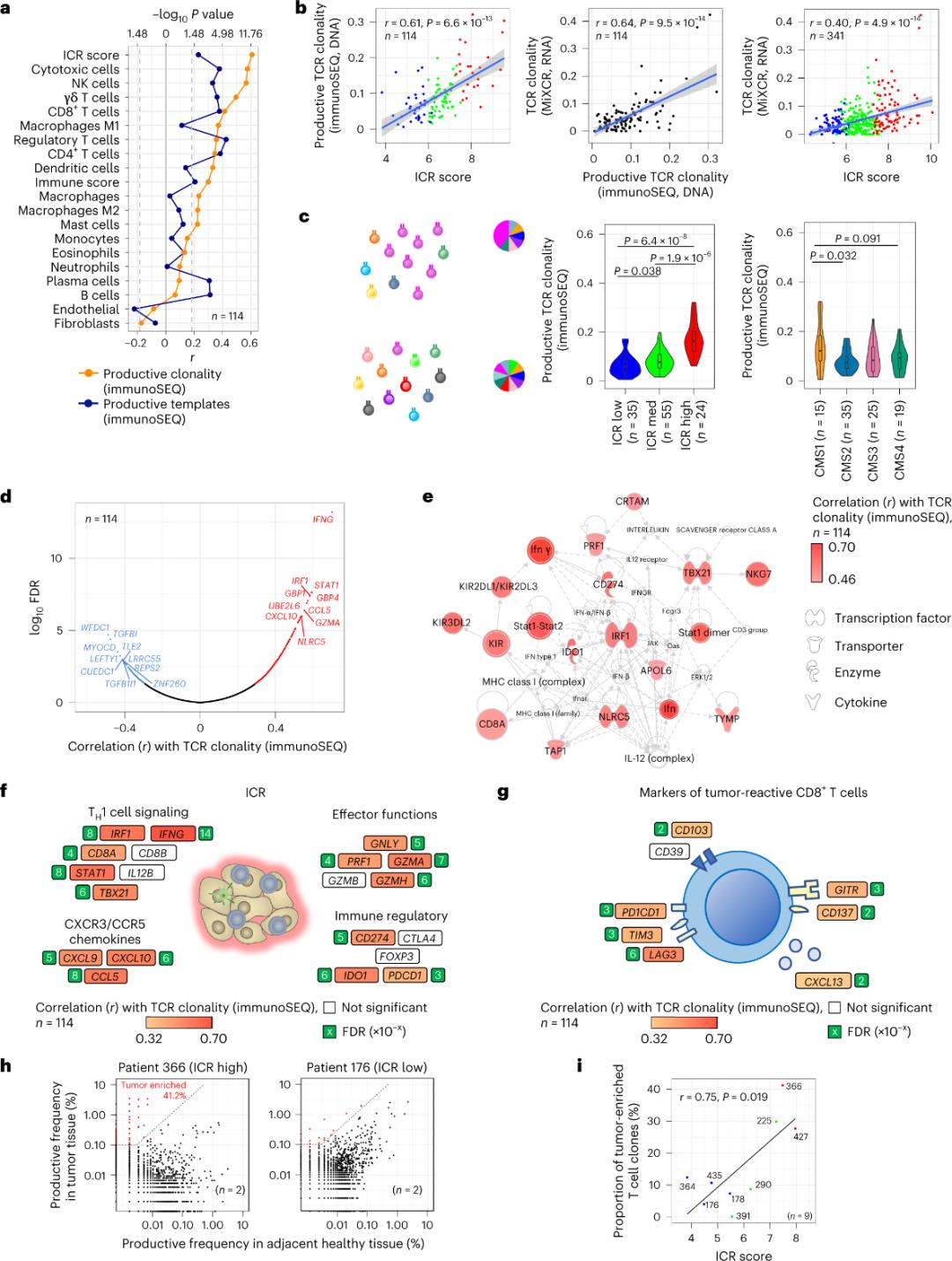
Figura 2. Mensurae TCR et correlatio cum genis immuno-relatis, subtypis immunibus et molecularibus.
Compositio microbiomatis in textibus sanis et carcinomatis coli
Investigatores sequentiationem 16S rRNA perfecerunt, DNA ex tumore congruenti et textu coli sano a 246 aegrotis extracto utentes (Figura 3a). Ad validationem, investigatores insuper data sequentiationis geni 16S rRNA ex 42 exemplaribus tumoris additis, quae DNA normale congruens ad analysin non habebant, analysaron. Primo, investigatores abundantiam relativam florae inter tumores congruentes et textum coli sanum comparaverunt. *Clostridium perfringens* significanter auctum est in tumoribus comparatum cum exemplaribus sanis (Figura 3a-3d). Nulla differentia significativa in diversitate alpha (diversitas et abundantia specierum in uno exemplo) inter exempla tumoris et sana erat, et modesta reductio in diversitate microbica observata est in tumoribus ICR-alto respectu tumorum ICR-humili.
Ad nexus clinicos inter perfiles microbiales et exitus clinicos detegendos, investigatores proposuerunt ut notitias sequentiationis geni 16S rRNA adhiberent ad proprietates microbiomatis quae superviventiam praedicunt identificandas. Apud AC-ICAM246, investigatores exemplar regressionis OS Cox perfecerunt quod 41 proprietates cum coefficientibus non nullis (cum periculo mortalitatis differentiali coniunctas) selectavit, classificatores MBR appellatos (Figura 3f).
In hac cohorte exercitationis (ICAM246), gradus MBR humilis (MBR<0, MBR humilis) cum periculo mortis significanter minore (85%) coniunctus est. Investigatores coniunctionem inter MBR humilem (periculum) et OS prolongatam in duabus cohortibus independenter validatis (ICAM42 et TCGA-COAD) confirmaverunt. (Figura 3) Studium correlationem fortem inter coccos endogastricos et gradus MBR ostendit, qui similes erant in tumore et in textu coli sano.
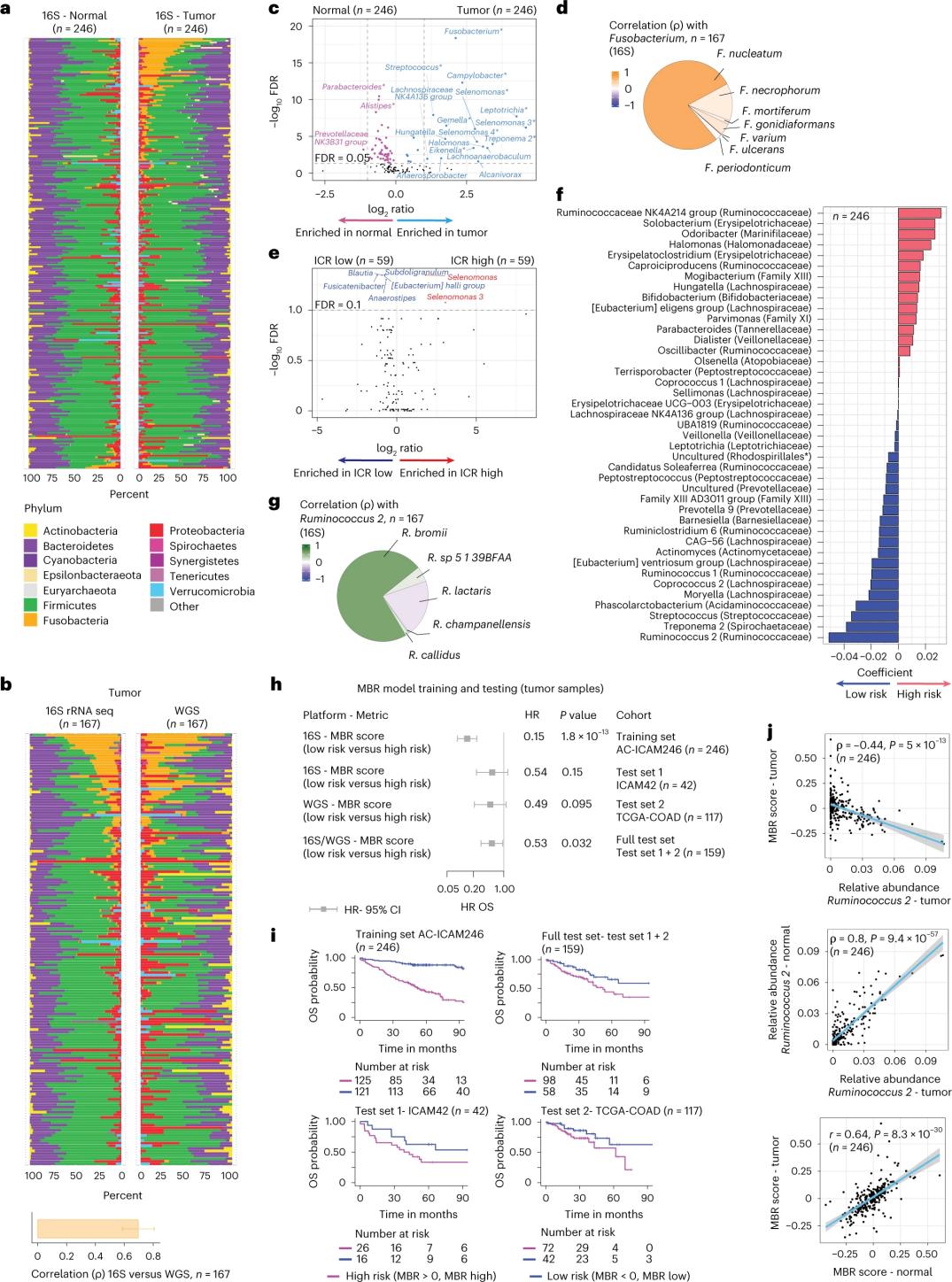
Figura 3. Microbioma in textibus tumoralibus et sanis et nexus cum ICR et superviventia aegrotorum.
Conclusio
Methodus multi-omica in hoc studio adhibita detectionem et analysin accuratam signaturae molecularis responsus immunis in cancro colorectali permittit et interactionem inter microbiome et systema immune revelat. Sequentiatio TCR profunda textuum tumoris et sanorum revelavit effectum prognosticum ICR fortasse ex facultate sua capiendi clones cellularum T tumore locupletatos et fortasse antigeno tumoris specificos oriri.
Per analysin compositionis microbiomis tumoris utens sequentiatione geni 16S rRNA in exemplaribus AC-ICAM, turma signaturam microbiomis (punctum periculi MBR) cum valido valore prognostico identificavit. Quamquam haec signatura ex exemplaribus tumoris derivata est, valida correlatio inter colorectum sanum et punctum periculi MBR tumoris exstabat, quod suggerit hanc signaturam compositionem microbiomis intestinalis aegrotorum capere posse. Coniungendo puncta ICR et MBR, possibile fuit bioindicem studentem multi-omicum identificare et validare qui superviventiam in aegrotis cum cancro coli praedicit. Collectio datorum multi-omica studii fontem praebet ad melius intellegendam biologiam cancri coli et ad adiuvandum inveniendas rationes therapeuticas personalizatas.
Tempus publicationis: Iun-XV-MMXXIII
 中文网站
中文网站